|
I am a Principal Researcher at Microsoft Research in Cambridge, MA.
I completed my PhD in Biophysics at Harvard University, where I worked with Sangeeta Bhatia at the Koch Institute for Integrative Cancer Research and was supported by the NSF Graduate Research Fellowship. I received my Bachelor of Science in Computer Science and Molecular Biology from MIT, where I was recognized as a Henry Ford II Scholar and with the AMITA Academic Award. Email / CV / Google Scholar / Twitter |

|
|
My research focuses on developing new AI methods to understand and design biology, with the ultimate aim of realizing precision biomedicines to improve human health. My work bridges the computational and experimental worlds to create new diagnostic and therapeutic biotechnologies and to achieve new insights into human disease. *Denotes co-first authorship. †Denotes corresponding authors. Formerly Ava Soleimany. |

|
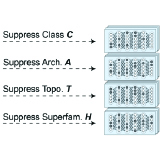
Sebastiano Cultrera di Montesano, Davide D'Ascenzo, Srivatsan Raghavan, Ava P. Amini, Peter S. Winter, Lorin Crawford Nature Computational Science, 2026 pdf / code We introduce a hierarchical cross-entropy loss for single-cell cell type annotation; this simple modification significantly improves out-of-distribution performance without added computational cost. |
|
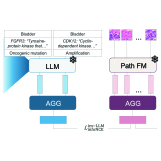
Ria Vinod, Ava P. Amini, Lorin Crawford, Kevin K. Yang PLOS Computational Biology, 2026 pdf / code We introduce trainable subnetworks to study how protein language models factorize structural knowledge among their learned parameters, revealing that even small changes in language modeling performance can significantly impair structure prediction. |
|
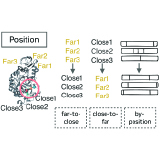
Ekaterina Redekop, Eric Zimmermann, Ava P. Amini, Alex X. Lu, Neil Tenenholtz, James B. Hall, Lorin Crawford, Kristen A. Severson arXiv, 2026 We develop MARBLE, a multimodal contrastive pretraining strategy that integrates structured biomarker knowledge into histopathology image representations, enabling data-efficient generalization to novel genomic biomarkers in precision oncology. |
|
Kieran Didi, Sarah Alamdari, Alex X. Lu, Bruce Wittmann, Kadina E. Johnston, Ava P. Amini, Ali Madani, Maya Czeneszew, Christian Dallago, Kevin K. Yang bioRxiv, 2026 pdf / website / code We present FLIP2, an expanded benchmark suite for protein fitness landscape prediction that introduces new datasets and generalization splits designed to rigorously evaluate machine learning models under real-world protein engineering scenarios. |
|
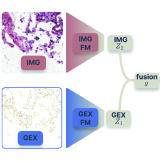
Till Richter, Eric Zimmermann, James Hall, Fabian J. Theis, Srivatsan Raghavan, Peter S. Winter, Ava P. Amini†, Lorin Crawford† bioRxiv, 2026 pdf / code We introduce a synergistic information score to evaluate multimodal cell foundation models, showing that alignment-based fusion is insufficient and that nonlinear, synergy-aware integration is required for genuinely complementary multimodal reasoning. |

|
Carmen Martin-Alonso*, Sarah Alamdari*, Tahoura S. Samad, Kevin K. Yang, Sangeeta N. Bhatia†, Ava P. Amini† Nature Communications, 2026 pdf / code / press We present CleaveNet, an end-to-end AI pipeline for the design of peptide-based protease substrates, enabling generation of peptides guided by a target cleavage profile for the design of efficient and selective substrates. |
|
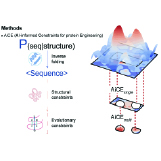
Kevin K. Yang, Ava P. Amini Cell, 2025 We discuss how the thoughtful application of existing deep learning models for fixed-backbone protein sequence design can effectively simplify and advance protein engineering. |
|
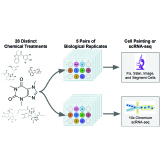
Joel Dapello, Marcel Nassar, Ridvan Eksi, Ban Wang, Jules Gagnon-Marchand, Kenneth T. Gao, Akram Baharlouei, Kyra Thrush-Evensen, Nina Riehs, Amy F. Peterson, Aniket Tolpadi, Abhejit Rajagopal, Henry E. Miller, Ashley Mae Conard, David Alvarez-Melis, Rory Stark, Simone Bianco, Morgan Levine, Ava P. Amini, Alex X. Lu, Nicolo Fusi, Ravi Pandya, Valentina Pedoia, Hana El-Samad NeurIPS, 2025 code We present scGeneScope, a large-scale dataset pairing single-cell RNA sequencing with Cell Painting microscopy images across 28 chemical treatments, alongside benchmarks for unimodal and multimodal treatment response modeling. |
|
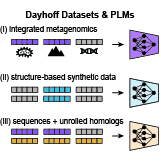
Kevin K. Yang*, Sarah Alamdari*, Alex J. Lee*, Kaeli Kaymak-Loveless, Samir Char, Garyk Brixi, Carles Domingo-Enrich, Chentong Wang, Suyue Lyu, Nicolo Fusi, Neil Tenenholtz†, Ava P. Amini† bioRxiv, 2025 pdf / code / press We introduce the Dayhoff Atlas, a centralized collection of protein sequence data and generative language models trained on GigaRef (3.34 billion sequences) and structure-based synthetic data, enabling improved protein generation and design. |
|
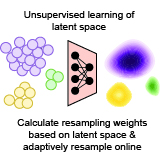
Zeinab Navidi, Akshaya Thoutam, Madeline Hughes, Srivatsan Raghavan, Peter S. Winter, Lorin Crawford†, Ava P. Amini† bioRxiv, 2025 pdf / code We present an adaptive resampling approach that addresses underrepresented cellular features in single-cell transcriptomics by dynamically resampling data based on learned latent structure during model training, improving downstream performance across tasks. |

|
Kasia Z. Kedzierska, Lorin Crawford, Ava P. Amini, Alex X. Lu Genome Biology, 2025 pdf / code We evaluate the performance of single-cell foundation models in ``zero-shot`` settings where they are used without any further training, and show that they are outperformed by simpler methods. |

|
Samir Char, Nathaniel Corley, Sarah Alamdari, Kevin K. Yang, Ava P. Amini† Bioinformatics, 2025 code ProtNote is a multimodal deep learning model that leverages free-form text to enable both supervised and zero-shot protein function prediction. |
|
|
Yemin Yu, Neil Tenenholtz, Lester Mackey, Ying Wei, David Alvarez-Melis, Ava P. Amini, Alex X. Lu arXiv, 2025 pdf / code We introduce a new deep learning framework, MICON, that models chemical compounds as treatments that induce counterfactual transformations of cell phenotypes for improved representation learning in microscopy-based morphological profiling. |

|
Alan DenAdel, Michelle L. Ramseier, Andrew W. Navia, Alex K. Shalek, Srivatsan Raghavan, Peter S. Winter, Ava P. Amini, Lorin Crawford American Journal of Human Genetics, 2025 code We develop a method for protecting against over-clustering in analysis of single-cell RNA sequencing data, by controlling for the impact of reusing the same data twice when performing differential expression analysis. |

|
Chentong Wang, Sarah Alamdari, Carles Domingo-Enrich, Ava P. Amini, Kevin K. Yang Current Opinion in Structural Biology, 2025 We review recent advances in deep generative models for protein design, with a particular focus on sequence-structure co-generation methods. |

|
Ajay Nadig†, Akshaya Thoutam, Madeline Hughes, Anay Gupta, Andrew W. Navia, Nicolo Fusi, Srivatsan Raghavan, Peter S. Winter, Ava P. Amini†, Lorin Crawford† bioRxiv, 2025 pdf / code We systematically investigate the consequences of training dataset composition on the behavior of deep learning models of single-cell transcriptomics, focusing on human hematopoiesis as a tractable model system and including cells from adult and developing tissues, disease states, and perturbation atlases. |
 
|
Kevin P. Greenman, Ava P. Amini†, Kevin K. Yang† PLOS Computational Biology, 2025 pdf / code We assess deep learning-based uncertainty quantification methods on protein sequence-function prediction tasks. |

|
Nuo Liu, Walaa E. Kattan, Benjamin E. Mead, Conner Kummerlowe, Thomas Cheng, Sarah Ingabire, Jaime H. Cheah, Christian K. Soule, Anita Vrcic, Jane K. McIninch, Sergio Triana, Manuel Guzman, Tyler T. Dao, Joshua M. Peters, Kristen E. Lowder, Lorin Crawford, Ava P. Amini, Paul C. Blainey, William C. Hahn, Brian Cleary, Bryan Bryson, Peter S. Winter, Srivatsan Raghavan, Alex K. Shalek Nature Biotechnology, 2024 We establish a method of pooling perturbations, like chemical compounds, followed by computational deconvolution to reduce required sample size, labor, and cost in high-throughput phenotypic screens. |

|
Alan DenAdel, Madeline Hughes, Akshaya Thoutam, Anay Gupta, Andrew W. Navia, Nicolo Fusi, Srivatsan Raghavan, Peter S. Winter, Ava P. Amini†, Lorin Crawford† bioRxiv, 2024 pdf / code We investigate the role of pre-training dataset size and diversity on the performance of single-cell foundation models on both zero-shot and fine-tuned tasks, observing that current methods plateau in performance with pre-training datasets that are only a fraction of the full size. |

|
Rebecca Boiarsky, Nalini M. Singh, Alejandro Buendia, Ava P. Amini, Gad Getz, David Sontag Nature Machine Intelligence, 2024 pdf / code We take a deep dive into scBERT, one recently developed transformer model for single-cell RNA-sequencing data, to develop a deeper understanding of the potential benefits and limitations of single-cell foundation models. |

|
Sarah Alamdari, Nitya Thakkar, Rianne van den Berg, Neil Tenenholtz, Robert Strome, Alan M. Moses, Alex X. Lu, Nicolo Fusi, Ava P. Amini†, Kevin K. Yang† bioRxiv, 2024 pdf / code We develop EvoDiff, a general-purpose discrete diffusion over protein sequences, that combines evolutionary-scale data with the distinct conditioning capabilities of diffusion models for controllable protein generation in sequence space. |

|
Peter S. Winter*, Michelle L. Ramseier*, Andrew W. Navia*, Sachit Saksena, Haley Strouf, Nezha Senhaji, Alan DenAdel, Mahnoor Mirza, Hyun Hwan An, Laura Bilal, Peter Dennis, Catherine S. Leahy, Kay Shigemori, Jennyfer Galves-Reyes, Ye Zhang, Foster Powers, Nolawit Mulugeta, Alejandro J. Gupta, Nicholas Calistri, Alex Van Scoyk, Kristen Jones, Huiyun Liu, Kristen E. Stevenson, Siyang Ren, Marlise R. Luskin, Charles P. Couturier, Ava P. Amini, Srivatsan Raghavan, Robert J. Kimmerling, Mark M. Stevens, Lorin Crawford, David M. Weinstock, Scott R. Manalis, Alex K. Shalek, Mark A. Murakami bioRxiv, 2024 Utilizing patient-derived xenograft (PDX) models and clinical trial specimens of acute lymphoblastic leukemia (ALL), we examined how genetic and transcriptional features co-evolve to drive progression during prolonged tyrosine kinase inhibitor response, uncovering a landscape of cooperative mutational and transcriptional escape mechanisms that differ from those causing resistance to first generation inhibitors. |

|
Francesca-Zhoufan Li, Ava P. Amini, Yisong Yue, Kevin K. Yang, Alex X. Lu International Conference on Machine Learning, 2024 To understand how the features learned in pretraining protein language models (PLMs) relate to and are useful for downstream tasks, we perform a systematic analysis of transfer learning using PLMs, conducting 370 experiments across a comprehensive suite of factors including different downstream tasks, architectures, model sizes, model depths, and pretraining time. |

|
Chibuikem Nwizu, Madeline Hughes, Michelle L. Ramseier, Andrew W. Navia, Alex K. Shalek, Nicolo Fusi, Srivatsan Raghavan, Peter S. Winter, Ava P. Amini†, Lorin Crawford† bioRxiv, 2024 We developed a non-parametric infinite mixture model that leverages Bayesian sparse priors to identify marker genes while simultaneously performing clustering on single-cell expression data. |

|
Kevin E. Wu, Kevin K. Yang, Rianne van den Berg, Sarah Alamdari, James Y. Zou, Alex X. Lu, Ava P. Amini† Nature Communications, 2024 pdf / code We present FoldingDiff, a diffusion-based generative model that generates protein backbone structures via a procedure inspired by the natural folding process. |
 
|
Carmen Martin-Alonso*, Shervin Tabrizi*, Kan Xiong*, Timothy Blewett, Sainetra Sridhar, Andjela Crnjac, Sahil Patel, Zhenyi An, Ahmet Bekdemir, Douglas Shea, Shih-Ting Wang, Sergio Rodriguez-Aponte, Christopher A. Naranjo, Justin Rhoades, Jesse D. Kirkpatrick, Heather E. Fleming, Ava P. Amini, Todd R. Golub, J. Christopher Love†, Sangeeta N. Bhatia†, Viktor A. Adalsteinsson† Science, 2024 pdf / MIT press / general press We develop intravenous priming agents that are given prior to a blood draw to increase the abundance of cell-free DNA in circulation, improving the sensitivity of liquid biopsy cancer diagnostic assays. |
 
|
Carolina Rios-Martinez, Nicholas Bhattacharya, Ava P. Amini, Lorin Crawford, Kevin K. Yang PLOS Computational Biology, 2023 We develop a self-supervised masked language model for biosynthetic gene clusters in bacteria, and leverage it for natural product classification. |

|
Ava P. Amini, Kevin K. Yang Nature Computational Science, 2023 A commentary on a recent image-inspired, diffusion model to generate new protein structures. |
 
|
Aereas Aung, Ang Cui, Laura Maiorino, Ava P. Amini, Justin R. Gregory, Maurice Bukenya, Yiming Zhang, Heya Lee, Christopher A. Cottrell, Duncan M. Morgan, Murillo Silva, Heikyung Suh, Jesse D. Kirkpatrick, Parastoo Amlashi, Tanaka Remba, Leah M. Froehle, Shuhao Xiao, Wuhbet Abraham, Josetta Adams, J. Christopher Love, Phillip Huyett, Douglas S. Kwon, Nir Hacohen, William R. Schief, Sangeeta N. Bhatia, Darrell J. Irvine Science, 2023 pdf / MIT press / general press We discover "sanctuaries" within lymph nodes that contain low proteolytic activity and act as a safe haven for vaccines, and demonstrate that this heterogeneity can be exploited to enhance vaccine-induced antibody response. |
 
|
Ava P. Amini*, Jesse D. Kirkpatrick*, Cathy S. Wang, Alex M. Jaeger, Susan Su, Santiago Naranjo, Qian Zhong, Tyler Jacks, Sangeeta N. Bhatia Nature Communications, 2022 pdf / supplement We engineer an integrated set of methods for measuring specific protease activities across the organismal, tissue, and cellular scales, and unify these methods into a methodological hierarchy that powers new biological insights into cancer. |
 
|
Ava P. Soleimany*, Carmen Martin-Alonso*, Melodi Anahtar*, Cathy S. Wang, Sangeeta N. Bhatia ACS Omega, 2022 pdf / code We build Protease Activity Analysis (PAA), a Python software package with a collection of data analytic and machine learning tools for analyzing protease activity data. |
 
|
Melodi Anahtar, Leslie W. Chan, Henry Ko, Aditya Rao, Ava P. Soleimany, Purvesh Khatri, Sangeeta N. Bhatia PNAS, 2022 pdf / code / press We develop a sensor-based, ML-driven system to diagnose pneumonia and classify its etiology, using machine learning to classify directly from molecular barcodes. |
 
|
Jesse D. Kirkpatrick, Ava P. Soleimany, Jaideep S. Dudani, Heng-Jia Liu, Hilaire C. Lam, Carmen Priolo, Elizabeth P. Henske, Sangeeta N. Bhatia European Respiratory Journal, 2022 We establish a sensor-based, ML-driven diagnostic for noninvasive, real-time monitoring of disease in a preclinical model of lymphangioleiomyomatosis (LAM), a rare lung disease. |
 
|
Ahmet Bekdemir, Eden E.L. Tanner, Jesse D. Kirkpatrick, Ava P. Soleimany, Samir Mitragotri, Sangeeta N. Bhatia Advanced Healthcare Materials, 2022 pdf / supplement We design an easily applicable, non-invasive formulation to deliver diagnostic nanosensors through the skin, enabling a sustained release diagnostic monitoring system for detecting thrombosis. |
 
|
Jiang He*, Lior Nissim*, Ava P. Soleimany*, Adina Binder-Nissim, Heather E. Fleming, Timothy K. Lu, Sangeeta N. Bhatia ACS Synthetic Biology, 2021 pdf / supplement We design a sense-and-respond system that integrates a synthetic gene circuit and nanotechnology detection tools for tumor-specific expression of heterologous biomarkers. |
 
|
Ava P. Soleimany*, Alexander Amini*, Samuel Goldman*, Daniela Rus, Sangeeta N. Bhatia, Connor W. Coley ACS Central Science, 2021 pdf / supplement A fast, scalable approach for uncertainty quantification in neural networks enables uncertainty-aware molecular property prediction, accelerated property optimization, and guided virtual screening. |
 
|
Ava P. Soleimany*, Jesse D. Kirkpatrick*, Susan Su, Jaideep S. Dudani, Qian Zhong, Ahmet Bekdemir, Sangeeta N. Bhatia Cancer Research, 2021 pdf / supplement We engineer a new class of enzyme activity probes that can be applied to fresh-frozen tissue sections to spatially localize protease activty, enabling new insights into the biology of protease dysregulation. |
 
|
Alexander Amini, Wilko Schwarting, Ava Soleimany, Daniela Rus NeurIPS, 2020 pdf / press We develop a novel algorithm for fast, scalable uncertainty quantification in highly complex, non-linear neural networks trained for regression tasks. |
 
|
Naveen K. Mehta, Roma V. Pradhan, Ava P. Soleimany, Kelly D. Moynihan, Adrienne M. Rothschilds, Noor Momin, Kavya Rakhra, Jordi Mata-Fink, Sangeeta N. Bhatia, K. Dane Wittrup, Darrell J. Irvine Nature Biomedical Engineering, 2020 pdf / supplement / press We optimize the immunogenicity of peptide-based antitumor vaccines in mice by tuning their pharmacokinetics via fusion of the peptide epitopes to protein carriers. |
 
|
Ava P. Soleimany, Sangeeta N. Bhatia Trends in Molecular Medicine, 2020 Review detailing how integrating techniques from multiple disciplines has developed engineered diagnostics that are selectively activated in disease states, highlighting their potential to realize the goals of precision medicine. |
 
|
Jesse D. Kirkpatrick*, Andrew D. Warren*, Ava P. Soleimany*, Peter M. K. Westcott, Justin C. Voog, Carmen Martin-Alonso, Heather E. Fleming, Tuomas Tammela, Tyler Jacks, Sangeeta N. Bhatia Science Translational Medicine, 2020 supplement / press / video We couple protease-responsive nanoparticle sensors with machine learning to engineer a sensitive and specific urinary test for lung cancer detection. |
 
|
Simone Schuerle, Maiko Furubayashi, Ava P. Soleimany, Tinotenda Gwisai, Wei Huang, Christopher A. Voigt, Sangeeta N. Bhatia ACS Synthetic Biology, 2020 supplement Magnetic nanoparticles that display genetically encoded targeting peptides to promote tumor accumulation and enhance MRI contrast. |
 
|
Colleen Loynachan*, Ava P. Soleimany*, Jaideep S. Dudani, Yiyang Lin, Adrian Najer, Ahmet Bekdemir, Qu Chen, Sangeeta N. Bhatia†, Molly M. Stevens† Nature Nanotechnology, 2019 supplement / data / press / video By leveraging the unique properties of catalytic nanomaterials, we develop a simple color-change urine test for detection of cancer in mice. |
 
|
Ava P. Soleimany, Harini Suresh, Jose Javier Gonzalez Ortiz, Divya Shanmugam, Nil Gural, John Guttag, Sangeeta N. Bhatia ICML Workshop on Computational Biology, 2019 Convolutional neural networks for automated segmentation and uncertainty estimation of microscopy images of malaria infection. |

|
Simone Schuerle, Ava P. Soleimany, Tiffany Yeh, Giridhar M. Anand, Moritz Haberli, Heather E. Fleming, Nima Mirkhani, Famin Qiu, Sabine Hauert, Xiaopu Wang, Bradley J. Nelson, Sangeeta N. Bhatia Science Advances, 2019 supplement / press / video Engineered microrobots that use magnetism to push drug-delivery nanoparticles out of blood vessels and into diseased tissue. |

|
Alexander Amini*, Ava P. Soleimany*, Wilko Schwarting, Sangeeta N. Bhatia, Daniela Rus AAAI/ACM Conference on Artificial Intelligence, Ethics, and Society, 2019, *Co-first authors MIT press / VentureBeat press Generalizable algorithm for mitigating hidden biases within training data, by leveraging learned latent distributions to adaptively re-weight the importance of certain data points while training. |
 
|
Alexander Amini, Ava Soleimany, Sertac Karaman, Daniela Rus, NeurIPS Workshop on Bayesian Deep Learning, 2017 Estimating uncertainty in neural networks for end-to-end control by exploiting feature map correlations during training. |
 
|
Nathaniel Roquet, Ava P. Soleimany, Alyssa C. Ferris, Scott Aaronson, Timothy K. Lu Science, 2016 supplement / press / blog Programming biological state machines that enable cells to remember and respond to a series of events. |
|
In addition to research, I am passionate about education and leadership, and strive to help and empower others to excel in their own pursuits. |
 
|
I am an organizer and lecturer for Introduction to Deep Learning (6.S191), MIT’s official introductory course on deep learning foundations and applications. Together with Alexander Amini, I have organized and developed all aspects of the course, including developing the curriculum, teaching the lectures, creating software labs, and collaborating with industry sponsors. All materials can be found online on the course website. |
 |
Co-founder, Momentum AI
I am a co-founder and director for Momentum AI, an outreach program that teaches AI and machine learning to under-resourced and under-served high school
students from the greater Boston area. Our two-week capstone program is a free, project-based deep dive into AI and is held on MIT's campus.
|
 |
Teaching Fellow, Harvard MCB294, Fall 2019, with Nancy Kleckner
|

|
Teaching Assistant, MIT 7.05, Spring 2016, with Matt Vander Heiden and Mike Yaffe
Teaching Assistant, MIT 7.05, Spring 2015, with Matt Vander Heiden and Mike Yaffe |
- [Nov. 2025] Modeling and targeting cell states in cancer, Machine Learning in Drug Discovery, Broad Institute.
- [Nov. 2025] Scaling generative models for functional protein design, Genetic Agency Technology Conference.
- [Feb. 2025] AI for Precision Health: Learning the language of nature and patients, Microsoft Research Forum.
- [Sep. 2024] Bridging biophysics and AI for generative protein design, MLCB Keynote.
- [Apr. 2024] Learning the language of biology: towards precision medicine, MIT IIA Summit.
- [Jan. 2024] How will AI transform precision medicine?, World Economic Forum.
- [Nov. 2023] Protein design with generative diffusion models, Broad Institute MIA Seminar.
- [Oct. 2022] Optimizing ex vivo organoids for precision cancer medicine, Microsoft Research Forum.
- [Oct. 2021] Leveraging uncertainty in machine learning to bridge computation and experimentation, Microsoft Research Forum.